HPC Basics
GRIT High Performance Computing Environment (hpc.grit.ucsb.edu)
The GRIT High Performance Computing (HPC) environment at the University of California, Santa Barbara provides researchers with a scalable, GPU-accelerated computing platform accessible through a web-based portal powered by Open OnDemand (OOD). The system is designed to support computational research workflows across domains including data science, machine learning, environmental modeling, and large-scale simulation.
The cluster is centrally accessed via hpc.grit.ucsb.edu, where researchers can launch interactive computing sessions, Jupyter notebooks, RStudio environments, and remote desktop sessions directly from a browser without requiring local software installation. Open OnDemand integrates with the cluster scheduler to provide fair-share resource allocation and secure multi-user access.
System Architecture
The GRIT cluster currently consists of ten compute servers running Ubuntu 24.04 LTS, providing a mix of high-memory nodes, CPU-optimized nodes, and GPU-accelerated nodes for machine learning and parallel workloads.
Key characteristics include:
-
Total CPU capacity: 664 CPU cores
-
Total system memory: approximately 7.5 TB RAM
-
GPU resources: NVIDIA L40S and A30 accelerators
-
Operating system: Ubuntu 24.04 LTS across all nodes
-
Access platform: Open OnDemand web portal
-
Scheduler: Slurm workload manager
Open OnDemand Research Portal
Researchers interact with the cluster through Open OnDemand, which provides:
-
browser-based JupyterLab, RStudio, and VS Code environments
-
interactive Linux desktop sessions for visualization and GUI applications
-
direct Slurm job submission and monitoring
-
secure multi-user authentication integrated with the university identity infrastructure
This interface significantly lowers the barrier to HPC adoption by eliminating the need for SSH configuration or command-line familiarity, enabling researchers and students to rapidly deploy computational workflows.
Research Applications
The GRIT HPC environment supports a broad set of research activities including:
-
machine learning and AI model training using NVIDIA GPUs
-
large-scale data analysis and statistical modeling
-
climate and environmental simulation
-
computational biology and genomics pipelines
-
interactive exploratory computing via Jupyter and RStudio
The combination of high-memory nodes, GPU accelerators, and an accessible web-based interface allows the platform to support both advanced HPC users and researchers new to large-scale computing.
Compute Resources
| Hostname | OS | RAM | CPU | GPU | RDP | Jupyterhub | R studio |
| hammer.eri.ucsb.edu | Ubuntu 24.04 |
128 GB | 44 Cores | No | Yes | Yes | Yes |
| anvil.eri.ucsb.edu | Ubuntu 24.04 |
384 GB | 24 Cores | No | Yes | Yes | No |
| forge.eri.ucsb.edu | Fedora 40 |
500 GB | 24 Cores | No | No | Yes | Yes |
| tong.eri.ucsb.edu | Fedora 41 |
256 GB | 32 Cores | No | No | No | No |
| bellows.eri.ucsb.edu | CentOS 8 |
1.5 TB | 96 Cores | No | No | No | No |
| biscuit.grit.ucsb.edu | Windows |
240 GB | 32 Cores | P100 | Yes | No | No |
| whpc-01.grit.ucsb.edu | Windows |
256GB | 32 Cores | L40S | Yes | No | No |
|
hpc.grit.ucsb.edu Cluster that contains the following servers |
Ubuntu 24.04 |
Yes | Yes | Yes | |||
| hpc-01.grit.ucsb.edu | Ubuntu 24.04 |
1.5 TB | 64 Cores | A30 (1) | OOD | OOD | OOD |
| hpc-02.grit.ucsb.edu | Ubuntu 24.04 |
1 TB | 48 Cores | L40S (4) | OOD | OOD | OOD |
| hpc-03.grit.ucsb.edu | Ubuntu 24.04 |
512GB | 24 Cores | L40S (2) | OOD | OOD | OOD |
| hpc-04.grit.ucsb.edu | Ubuntu 24.04 |
32GB | 8 Cores | L40S (2) | No | No | No |
| hpc-05.grit.ucsb.edu | Ubuntu 24.04 |
512GB | 72 Cores | No | OOD | OOD | OOD |
| hpc-06.grit.ucsb.edu | Ubuntu 24.04 |
512GB | 72 Cores | No | OOD | OOD | OOD |
| hpc-07.grit.ucsb.edu | Ubuntu 24.04 |
512GB | 72 Cores | No | OOD | OOD | OOD |
| hpc-08.grit.ucsb.edu | Ubuntu 24.04 |
512GB | 72 Cores | No | OOD | OOD | OOD |
| hpc-09.grit.ucsb.edu | Ubuntu 24.04 |
1TB | 144 Cores | No | OOD | OOD | OOD |
| hpc-10.grit.ucsb.edu | Ubuntu 24.04 |
1TB | 48 Cores | L40S | OOD | OOD | OOD |
| hpc-11.grit.ucsb.edu | Ubuntu 24.04 |
512GB | 48 Cores | L40S | OOD | OOD | OOD |
hpc.grit.ucsb.edu can be accessed via Open On Demand. More info here:
https://bookstack.grit.ucsb.edu/books/hpc-usage/chapter/openondemand-ood
Typically we set up some local scratch space for you to store your stuff.
#Info obtained from each system with: scontrol show node | grep CPU
Access
Access to the HPC systems using secure shell with your username and password from a command line is limited to campus IP addresses, and from other machines within the GRIT ecosystem. For example:
# connect to the bastion host
ssh username@ssh.grit.ucsb.edu
# or go straight to your HPC machine, e.g. bellows.eri.ucsb.edu
ssh username@bellows.grit.ucsb.edu
For much more detail, and some instructions on more efficient access see this page:
https://bookstack.grit.ucsb.edu/books/remote-access/page/ssh-key-setup
This page has information for connecting from off campus:
https://bookstack.grit.ucsb.edu/books/remote-access/page/ssh-mfa-setup
Storage Notes
Almost all storage in GRIT is available to all systems that you can access, this includes your home directory. For example /home/<user> will be the same on all systems, as will /home/<lab>. The benefit of this is it makes moving between machines very easy. The downside is that writing to these traverses the network, which can be an issue for jobs writing or reading lots of data.
Each of the HPC machines has local scratch storage (meaning locally mounted to the computer and *not* backed up). On bellows it's under /scratch (so, scratch/<lab>/ for example), on some of the older machines it would be/raid/scratch/. This is specifically used for reading and writing by jobs running on the machine, available as a community resource and not for long term storage.
Moving Data
rsync is a very powerful and widely used tool for moving data. The manual page has many useful examples (from the command line type "man rsync"). Here are a couple of examples to get you started:
# the command format is #rsync # # So the following copies from a local folder to a destination folder on a remote host named bellows.eri.ucsb.edu: rsync -avr /data/ username@bellows.eri.ucsb.edu:/some/other/folder/ # the -avr switches are: # 'a' for archive mode (when in doubt use this) # 'v' for verbose (rsync will tell you what is going on) # 'r' for recursive, recurse into directories
One trick to learn with rsync is the difference between leaving the trailing slash on or off.
# this command copies contents of /data/ to the destination directory /some/other/folder/ rsync -avr /data/ username@bellows.eri.ucsb.edu:/some/other/folder/ # ... while this command creates a folder 'data' on the destination and copies all of its contents: rsync -avr /data username@bellows.eri.ucsb.edu:/some/other/folder/
When in doubt, test with --dry-run, and rsync will tell you what would have happened:
rsync -avr --dry-run /data username@bellows.eri.ucsb.edu:/some/other/folder/
Running Code
To run your code and use the HPC machine in a fair and efficient way, you'll use a queuing system. See: https://bookstack.grit.ucsb.edu/books/hpc-usage/page/slurm-usage
A few key slurm commands are:
# submit a job (where slurm_test.sh is a shell script for invoking a program) sbatch slurm_test.sh # show the whole queue squeue -a # look at a job's details scontrol show job
screen can be used when running jobs to allow you to disconnect your computer from a remote terminal session, for example when running a very long rsync job. See: https://bookstack.grit.ucsb.edu/books/hpc-usage/page/the-screen-program
Other Notes
All the HPC systems are built on the ZFS file system. To see information, for example e.g. about how much space is available:
[username@hpcsystem ~]$ zfs list NAME USED AVAIL REFER MOUNTPOINT sandbox1 21.3T 4.00T 96K /mnt/sandbox1 sandbox2 21.3T 4.00T 96K /mnt/sandbox2 sandbox3 21.3T 4.00T 21.1T /mnt/sandbox3
... this system has 4 Terabytes available for storage.
# Commands for retrieving the system specifications (OS, RAM, Cores): cat /etc/redhat-release free -h # get number of cpu's for slurm scontrol show node | grep CPU # old way: grep 'processor' /proc/cpuinfo | uniq
Jupyterhub (SLURM)
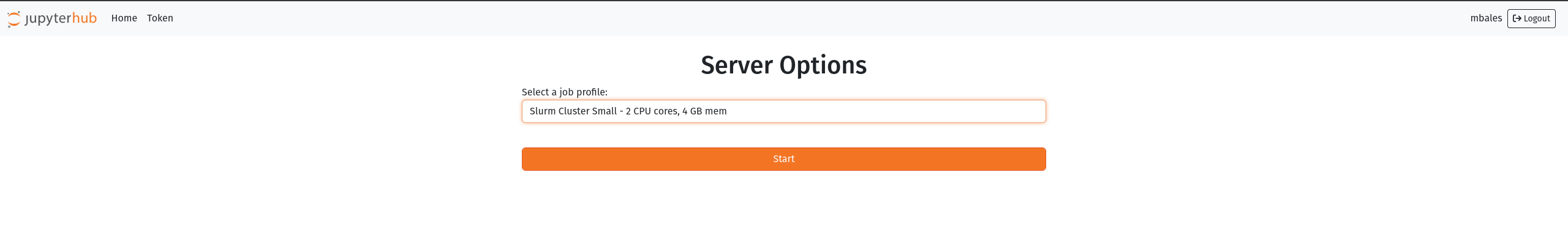
we are moving toward integrating jupyterhub with the SLURM work queuing system. What this means practically is we will be able to better manage system resource usage to improve system stability and accessibility. Upon login you will be presented with a profile selection screen:
Clicking the profile dropdown will present other options if you need more resources. Note that if you the system does not have the available resources your instance will be placed in the queue and spawn when resources are available. Please check the zabbix HPC page here: https://zabbix.grit.ucsb.edu/zabbix.php?action=dashboard.view&dashboardid=189&from=now%2Fd&to=now to get an idea of the current slurm queue and resources in use. After selecting your profile and clicking start it may take up to 1 minute for the instance to spawn.
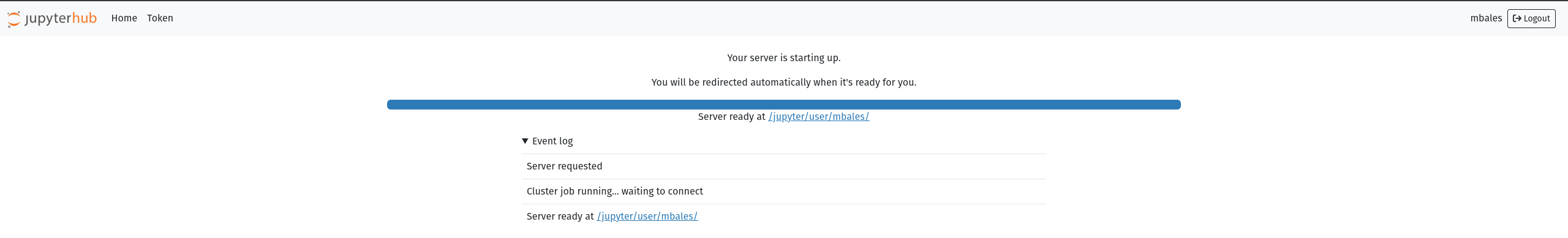
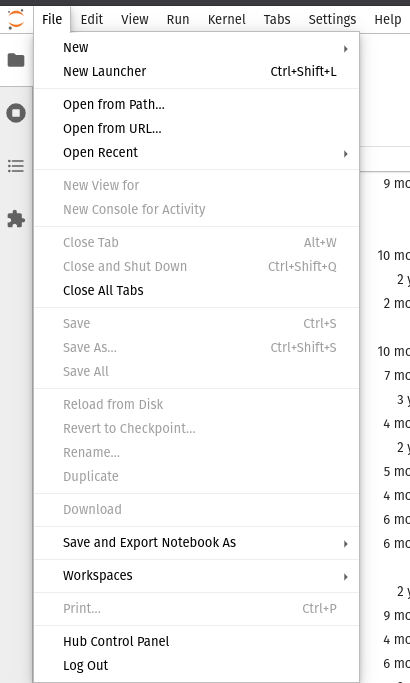
You will be automatically redirected when your instance is ready. Note that your instance is persistent until you manually kill your jupyterhub instance from the File drop down menu as shown below:
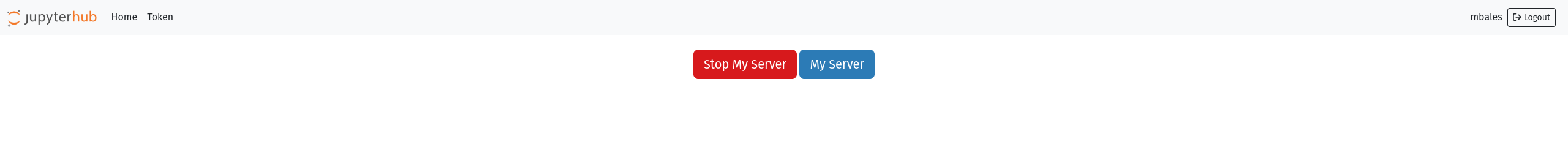
Select the Hub Control Panel option, this will open a new tab with the following options:
Selecting Stop My Server will kill your jupyterhub instance and remove the job from the SLURM queue.